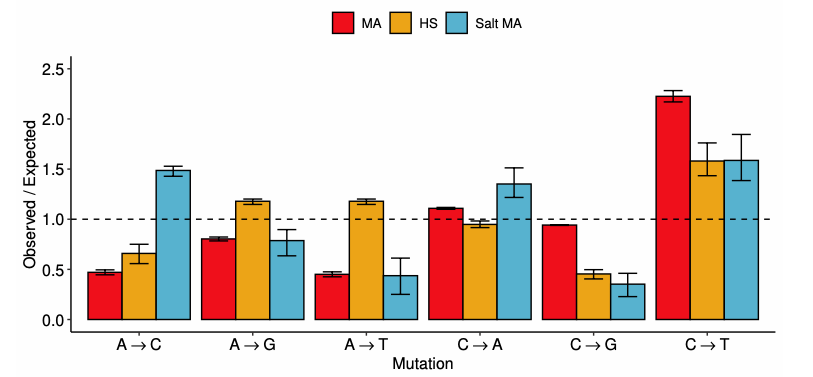
The genetic basis of adaptation is driven not only by selection, but also by the spectrum of available mutations. Given that the rate of mutation is not uniformly distributed across the genome and varies depending on the environment, understanding the signatures of selection across the genome is aided by first establishing what the expectations of genetic change are from mutation. To determine the interaction between salt stress, selection, and mutation across the genome, we compared the rates and patterns of mutation observed in a selection experiment for salt tolerance in Chlamydomonas reinhardtii to those observed in mutation accumulation experiments with and without salt exposure. We found that salt stress led to an increased rate of indel mutations, but that many of these mutations were removed under selection. Finally, lines adapted to salt also showed excess clustering of mutations in the genome and the co-expression network, suggesting a role for positive selection in retaining mutations in particular compartments of the genome during the evolution of salt tolerance. This study shows that characterizing mutation rates and spectra expected under stress helps disentangle the effects of environment and selection during adaptation.
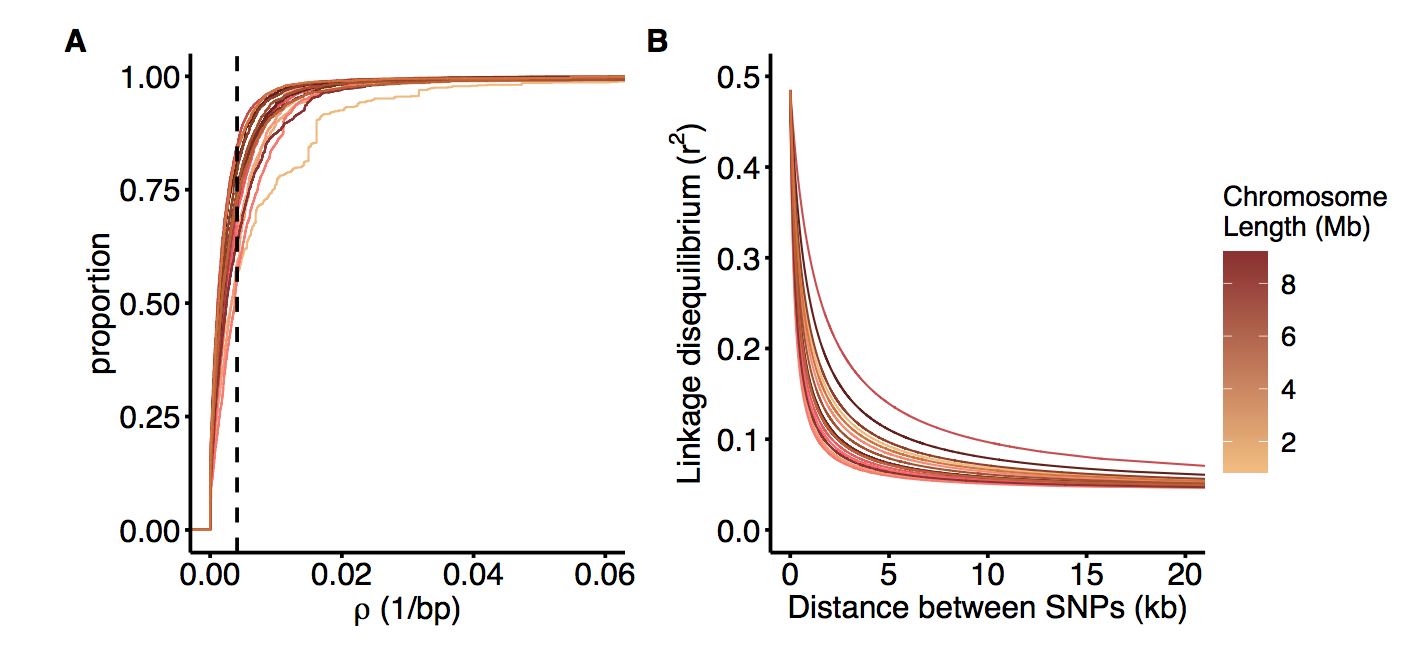
Recombination confers a major evolutionary advantage by breaking up linkage disequilibrium between harmful and beneficial mutations, thereby facilitating selection. However, in species that are only periodically sexual, such as many microbial eukaryotes, the realized rate of recombination is also affected by the frequency of sex, meaning that infrequent sex can increase the effects of selection at linked sites despite high recombination rates. Despite this, the rate of sex of most facultatively sexual species is unknown. Here, we use genomewide patterns of linkage disequilibrium to infer fine-scale recombination rate variation in the genome of the facultatively sexual green alga Chlamydomonas reinhardtii. We observe recombination rate variation of up to two orders of magnitude and find evidence of recombination hotspots across the genome. Recombination rate is highest flanking genes, consistent with trends observed in other nonmammalian organisms, though intergenic recombination rates vary by intergenic tract length. We also find a positive relationship between nucleotide diversity and physical recombination rate, suggesting a widespread influence of selection at linked sites in the genome. Finally, we use estimates of the effective rate of recombination to calculate the rate of sex that occurs in natural populations, estimating a sexual cycle roughly every 840 generations. We argue that the relatively infrequent rate of sex and large effective population size creates a population genetic environment that increases the influence of selection on linked sites across the genome.
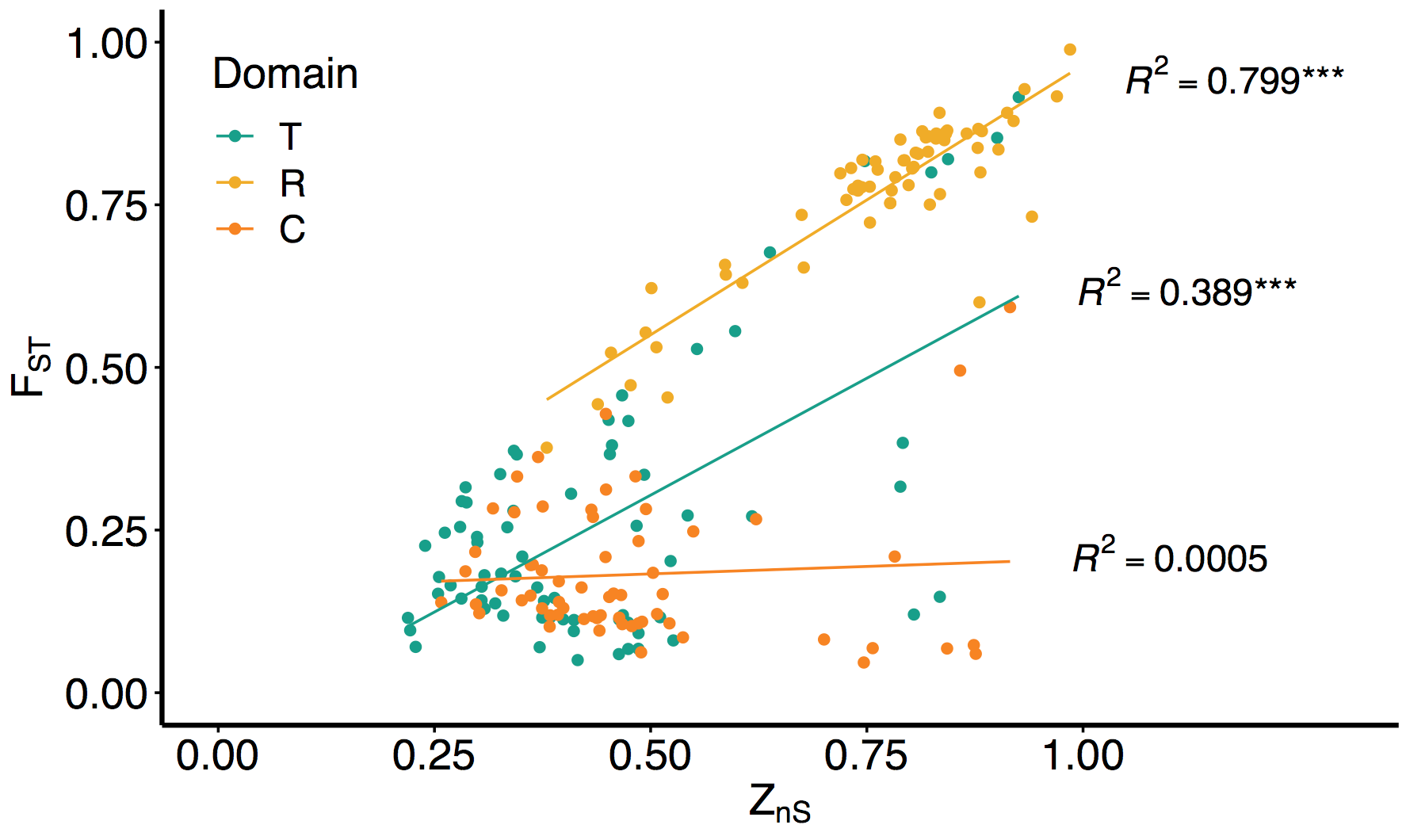
‘Sex’ in C. reinhardtii is determined by a dimorphic, crossover-suppressed mating type locus, which encodes an individual as either MT+ or MT-. C. reinhardtii is isogamous (equal sized gametes) which means there are no sex differences; hence the term ‘mating type’. The two alleles of the MT locus can be thought of as ‘X and Y chromosomes’, except that each individual only carries one and that the alleles are genetically very similar.
Unexpectedly similar, that is. Work has shown that the two MT alleles of C. reinhardtii are hardly differentiated from one another. Meanwhile, its multicellular, anisogamous relative Volvox carteri has shown over 100x more differentiation between its own MT alleles (Ferris 2010). Later work showed evidence for gene conversion, a form of noncrossover recombination, in two C. reinhardtii genes that had copies in both MT alleles, suggesting that genetic exchange was preventing the two alleles from diverging.
For this project, we estimate levels of genetic exchange across the MT locus of C. reinhardtii, and undertake a population genetic analysis to find out whether gene conversion is sufficient to avoid the differentiation and degeneration that is often seen in sex determining loci, asking the questions:
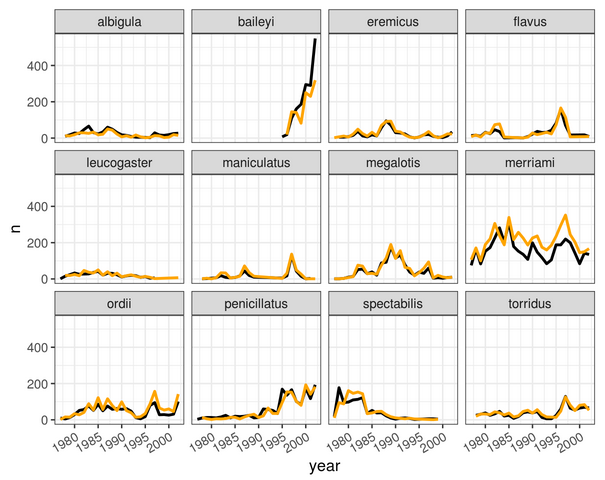
During the Fall 2018 and Fall 2019 semesters, I was part of the instructor team for a third-year course on quantitative methods in R, pitched by UofT Coders and adapted from Dr. Christie Bahlai’s Reproducible Quantitative Methods course. The course aims to teach undergraduate students at the third-year level or above the foundations for reproducible data analysis in R, and is taught entirely using the live-coding format used in Carpentries workshops. The course material is openly available on the course website.

As part of my undergraduate research work, I attempted to identify endophytic microbes that may potentially be used as biocontrols against the Côte d’Ivoire Lethal Yellowing (CILY) disease, which had previously destroyed approximately 350 ha of coconut trees in the West African country. This was done by creating profiles of the microbial communities present within both infected and uninfected trees. A ‘dilution-to-extinction’ method of high-throughput culturing was used in order to cultivate both bacterial and fungal endophytes directly from samples collected from the town of Grand-Lahou, followed by molecular analyses for strain identification. Our results identified Trichoderma, Penicillium, Bacillus, and Pseudomonas as endophytes present within coconut palms. Several of these endophytes have historically been used as part of microbial biocontrol consortia against other plant pathogens such as Fusarium, and show promise as potential biocontrols to be tested for use against CILY in field studies.
Related publication: Morales-Lizcano et al. (2017) Microbial diversity in leaves, trunk and rhizosphere of coconut palms (Cocos nucifera L.) associated with the coconut lethal yellowing phytoplasma in Grand-Lahou, Côte d’Ivoire. Available online.
Bonus: Here’s a fun and largely jargon-free one-minute video summary of the project that I made.